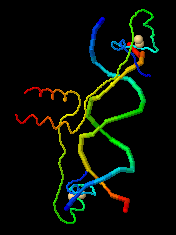Each of the views in the next three tables is achieved with
one click of the
mouse.
When actually using
FirstGlance, the molecular
views are
much larger .
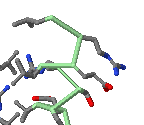
 Initial view (Cartoon plus Ligands+)
of Gal4 transcriptional
regulator (DNA-binding domain only) bound
to DNA
(1D66).
Each chain is a different color.
Initial view (Cartoon plus Ligands+)
of Gal4 transcriptional
regulator (DNA-binding domain only) bound
to DNA
(1D66).
Each chain is a different color.
|
 Secondary Structure
view of Gal4 with alpha helices
as "rockets"
(1D66).
Compare with another molecule below showing
a beta sheet.
Secondary Structure
view of Gal4 with alpha helices
as "rockets"
(1D66).
Compare with another molecule below showing
a beta sheet.
|
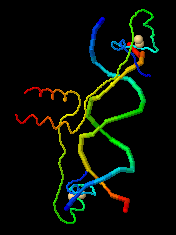
 N->C Rainbow view of Gal4:DNA showing
Amino/5'
and
Carboxy/3'
termini.
(1D66).
N->C Rainbow view of Gal4:DNA showing
Amino/5'
and
Carboxy/3'
termini.
(1D66).
|
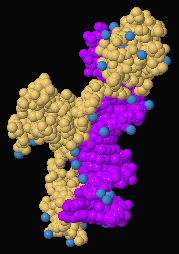 Composition view of Gal4:DNA indicating
Protein,
DNA,
and
Water
(1D66).
Composition view of Gal4:DNA indicating
Protein,
DNA,
and
Water
(1D66).
|
 Hydrophobic/Polar view of Gal4:DNA indicating
Hydrophobic
and
Polar residues on the surface.
(1D66).
Hydrophobic/Polar view of Gal4:DNA indicating
Hydrophobic
and
Polar residues on the surface.
(1D66).
|
 Charge view of Gal4:DNA indicating
Hydrophobic,
Polar uncharged,
Cationic (+),
Anionic (-),
and
Backbone atoms
on the surface.
(1D66).
Charge view of Gal4:DNA indicating
Hydrophobic,
Polar uncharged,
Cationic (+),
Anionic (-),
and
Backbone atoms
on the surface.
(1D66).
|
Atomic detail is available in Vines.. views.
The Slab.. option (not pictured) hides
atoms distant from the point of interest.
(1D66).
|
 Secondary Structure view of Streptococcal
protein G
(1PDB)
indicating
beta-strands,
alpha-helices
and
turns.
Secondary Structure view of Streptococcal
protein G
(1PDB)
indicating
beta-strands,
alpha-helices
and
turns.
|
Begin Contacts.. Section
The
Contacts.. tool of
FirstGlance produced
the following views.
The snapshots below are much smaller than
the actual views when using
FirstGlance.
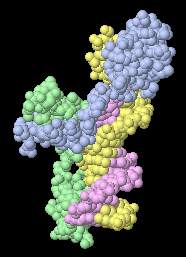 Contacts.. enables you to see how the chains
(each a different color) touch each other.
(View obtained with 1 click.)
Optionally, you can select a target (by clicking on chains, groups, or atoms)
in order to display the putative non-covalent bonds to the target.
(1D66)
Contacts.. enables you to see how the chains
(each a different color) touch each other.
(View obtained with 1 click.)
Optionally, you can select a target (by clicking on chains, groups, or atoms)
in order to display the putative non-covalent bonds to the target.
(1D66)
|
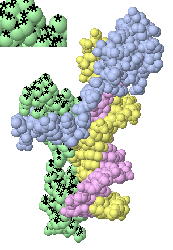 Clicking on one protein chain (green) marks it as the target
with a black asterisk (magnified in inset) on each atom.
(View of
1D66
obtained with 2 clicks.)
Clicking on one protein chain (green) marks it as the target
with a black asterisk (magnified in inset) on each atom.
(View of
1D66
obtained with 2 clicks.)
|
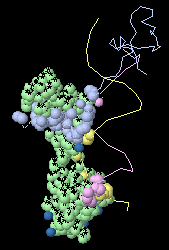 Only the atoms that contact the green target chain
are now shown.
(View obtained with 3 clicks;
1D66).
These contacts can be shown in great
detail, in seven subsets of different kinds of
non-covalent bonds, as shown below.
Only the atoms that contact the green target chain
are now shown.
(View obtained with 3 clicks;
1D66).
These contacts can be shown in great
detail, in seven subsets of different kinds of
non-covalent bonds, as shown below.
|
|
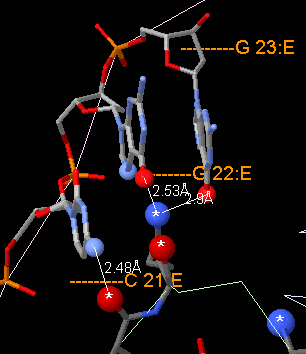
Contacts 1: non-water hydrogen bonds.
(Definition.)
At left,
target Gal4 protein
atoms (larger balls, bottom, marked with white asterisks)
are shown within hydrogen-bonding
distances of DNA atoms (smaller balls, top).
These hydrogen bonds are to a specific sequence of bases.
The DNA sequence
recognized by Gal4, CGG (in chain E), can be visualized here.
Atoms represented as sticks are too distant from the protein to be
hydrogen-bonded, but are shown for context.
(View of
1D66
obtained with 6 clicks, plus additional double-clicks
for each distance measurement.)
Double-clicking atoms reports distances or angles.
More..
In this view (and many views below), atoms are colored by element:
|
 Contacts 2 view of 1D66:
putative hydrogen bonds to
water.
(Definition.)
Contacts 2 view of 1D66:
putative hydrogen bonds to
water.
(Definition.)
|
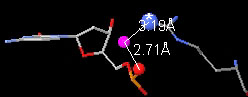 Contacts 3 view of 1D66:
putative
water
bridges, a subset of hydrogen bonds to water.
One putative bridge from 1D66 is shown above.
(Definition.)
Contacts 3 view of 1D66:
putative
water
bridges, a subset of hydrogen bonds to water.
One putative bridge from 1D66 is shown above.
(Definition.)
|
 Clicking on any atom reports its identity to the lower left
of the molecule, as shown above. ("NH2" means Nitrogen, Eta2.
More..)
Clicking on any atom reports its identity to the lower left
of the molecule, as shown above. ("NH2" means Nitrogen, Eta2.
More..)
|
 Contacts 4 view of 1D66:
van der Waals (hydrophobic) interactions.
(Definition.)
Here, the target chain atoms are spacefilled to van der Waals
radii, while the contacting atoms are small balls.
Contacts 4 view of 1D66:
van der Waals (hydrophobic) interactions.
(Definition.)
Here, the target chain atoms are spacefilled to van der Waals
radii, while the contacting atoms are small balls.
|
 Contacts 5: protein salt bridges.
(Definition.)
Contacts 5: protein salt bridges.
(Definition.)
|
 Contacts 6 view of 1BL8:
protein cation-pi orbital interactions.
(Definition.)
Contacts 6 view of 1BL8:
protein cation-pi orbital interactions.
(Definition.)
|

|
Contacts 7: metal and misc. interactions.
Here the interactions of
iron
in heme are shown (from
2HHD).
(Definitions.)
|
Begin Hide.., Find.. Section
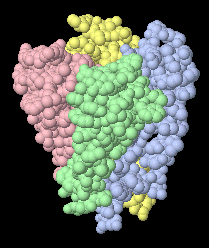 This potassium channel
(1BL8)
is a homotetramer (four identical chains).
This view is obtained with one click on Contacts...
This potassium channel
(1BL8)
is a homotetramer (four identical chains).
This view is obtained with one click on Contacts...
|
 Here, the Hide.. dialog of FirstGlance has been
used to hide the front two chains (3 more clicks). Now we can see three potassium
ions, and one water molecule.
(View of 1BL8).
Here, the Hide.. dialog of FirstGlance has been
used to hide the front two chains (3 more clicks). Now we can see three potassium
ions, and one water molecule.
(View of 1BL8).
|
 Hydrophobic/Polar shows that most of the surface of this
molecule is hydrophobic (gray), consistent with its transmembrane
location.
(View of
1BL8
obtained with 1 click.)
Compare with the
same view for Gal4.
Hydrophobic/Polar shows that most of the surface of this
molecule is hydrophobic (gray), consistent with its transmembrane
location.
(View of
1BL8
obtained with 1 click.)
Compare with the
same view for Gal4.
|
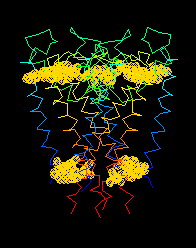 The Find.. dialog of FirstGlance was
used to locate all tryptophan residues (yellow clouds). Clearly,
they sit at the lipid-water boundaries.
(View of
1BL8
obtained with 2 clicks plus typing "TRP".)
The Find.. dialog of FirstGlance was
used to locate all tryptophan residues (yellow clouds). Clearly,
they sit at the lipid-water boundaries.
(View of
1BL8
obtained with 2 clicks plus typing "TRP".)
|
Begin Structural Bioinformatics Resources Section
FirstGlance has numerous links to structural bioinformatics
servers and resources.
One of the most important is a tool for obtaining
the
Biological Unit (functional assembly), the MakeMultimer server by Michael Palmer.
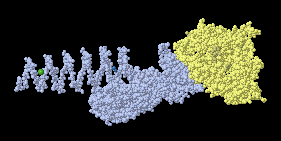 Here is
1K28
Notice the implausible folding of the left end,
which suggests that some of the chains are missing in this model,
(the asymmetric unit).
(View obtained with 1 click.)
Here is
1K28
Notice the implausible folding of the left end,
which suggests that some of the chains are missing in this model,
(the asymmetric unit).
(View obtained with 1 click.)
|
Might this molecule have formed specific oligomers, with more chains,
in the protein crystal? The answer is provided by the Biological Unit
section under the Resources Tab
|
 The Biological Unit is obtained with a few clicks, and reveals
that the molecule existed in the crystal as a hexamer,
rather than a dimer. The hexamer is formed from three dimers by symmetry
operations. A link back to FirstGlance at the MakeMultimer Server displays the above
specific oligomer.
The Biological Unit is obtained with a few clicks, and reveals
that the molecule existed in the crystal as a hexamer,
rather than a dimer. The hexamer is formed from three dimers by symmetry
operations. A link back to FirstGlance at the MakeMultimer Server displays the above
specific oligomer.
|
What is this amazing molecular machine? To find out,
view
1K28
in FirstGlance.
Crystallographic models often have portions missing -- regions where the protein in the crystal
was disordered ("blurry").
FirstGlance makes these obvious by highlighting areas with missing residues with
Empty Baskets (not yet shown here in a snapshot).
Nearly 50 amino acids are missing from the asymmetric unit of 1k28, (so nearly
150 from the specific hexamer)!
|
Begin More Views.. Section
 Within FirstGlance, More Views.. offers a display of all protein salt bridges
throughout the molecule. Here, all salt bridges in an SH3 domain
1B07
are shown. (View obtained with 2 clicks.)
Within FirstGlance, More Views.. offers a display of all protein salt bridges
throughout the molecule. Here, all salt bridges in an SH3 domain
1B07
are shown. (View obtained with 2 clicks.)
|
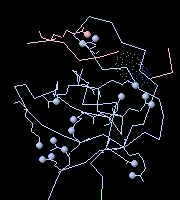 When the same view is colored by chain, it is easy to see
that one of the salt bridges is between two chains.
(View obtained with 1 additional click.)
When the same view is colored by chain, it is easy to see
that one of the salt bridges is between two chains.
(View obtained with 1 additional click.)
|
 The same dialog displays all protein cation-pi orbital interactions
throughout the molecule.
(View obtained with 2 clicks.)
The same dialog displays all protein cation-pi orbital interactions
throughout the molecule.
(View obtained with 2 clicks.)
|
 In the Views tab you can
color the molecule by Local Uncertainty (crystallographic "temperature" or "B factor").
The positions of the red atoms in Gal4:DNA model above have the least
certainty.
(View of
1D66
obtained with 3 clicks.)
In the Views tab you can
color the molecule by Local Uncertainty (crystallographic "temperature" or "B factor").
The positions of the red atoms in Gal4:DNA model above have the least
certainty.
(View of
1D66
obtained with 3 clicks.)
|